| Protein Superfamily | Index in Query Genome | Sequence Header | Detection Method |
|---|---|---|---|
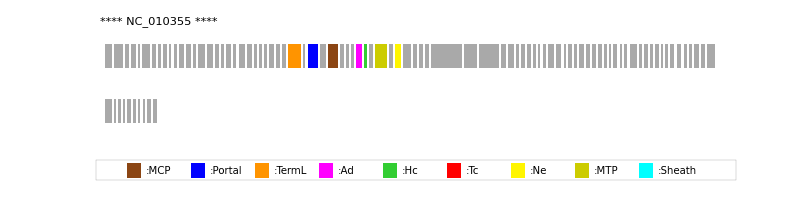
| MCP | 33 | lcl|NC_010355.1_cdsid_YP_001686874.1 | HHsearch (proba=100.00%) |
| Portal | 31 | lcl|NC_010355.1_cdsid_YP_001686872.1 | HHsearch (proba=100.00%) |
| TermL | 29 | lcl|NC_010355.1_cdsid_YP_001686870.1 | Blast (e-value=1e-40) |
| MTP | 40 | lcl|NC_010355.1_cdsid_YP_001686881.1 | Blast (e-value=4e-27) |
| Sheath | NOT FOUND | ||
| Ad1 | 37 | lcl|NC_010355.1_cdsid_YP_001686878.1 | HHsearch (proba=100.00%) |
| Hc1 | 38 | lcl|NC_010355.1_cdsid_YP_001686879.1 | HHsearch (proba=98.90%) |
| Ne1 | 42 | lcl|NC_010355.1_cdsid_YP_001686883.1 | HHsearch (proba=99.60%) |
| Tc1 | NOT FOUND | ||
| Ad2 | NOT FOUND | ||
| Hc2 | NOT FOUND | ||
| Tc2 | NOT FOUND | ||
| Ad3 | NOT FOUND | ||
| Hc3 | NOT FOUND | ||
| Ad4 | NOT FOUND |
| Proteins Superfamilies | Observed Intergene Distance | Expected Intergene Distance |
|---|---|---|
| Ne1<->Hc1 | 4 | 2 |
| Ne1<->Ad1 | 5 | 3 |
| Protein Superfamily | Corresponding Protein in Aclame | Aclame Phage | Seq. Identity (%) |
|---|---|---|---|
| MCP | protein:vir:4456 protein:vir:485 protein:vir:100247 | ST64B P27 Bcep176 | 39 % 39 % 35 % |
| Portal | protein:vir:80333 protein:vir:1431 protein:vir:100150 | phi644-2 phiE125 phi1026b | 31 % 31 % 30 % |
| TermL | protein:vir:100097 protein:vir:481 protein:vir:81178 | phi1026b P27 Geobacillus virus E2 | 25 % 25 % 24 % |
| MTP | protein:vir:97240 protein:vir:95133 protein:vir:80398 | M6 PA73 BcepGomr | 25 % 25 % 21 % |
| Ad1 | protein:vir:1435 protein:vir:80320 protein:vir:8430 | phiE125 phi644-2 Omega | 16 % 15 % 18 % |
| Hc1 | protein:vir:97237 protein:vir:80382 protein:vir:107669 | M6 BcepGomr T1 | 19 % 17 % 20 % |
| Ne1 | protein:vir:103280 protein:vir:94994 protein:vir:79638 | JK06 KS7 TLS | 23 % 20 % 19 % |