| Protein Superfamily | Index in Query Genome | Sequence Header | Detection Method |
|---|---|---|---|
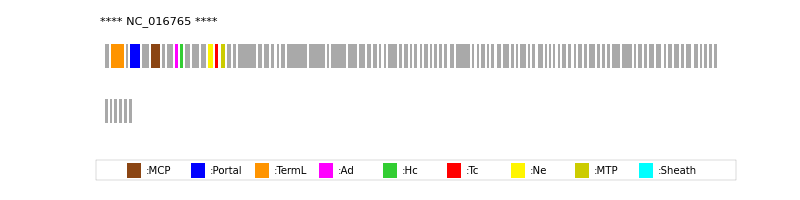
| MCP | 6 | lcl|NC_016765.1_cdsid_YP_005098209.1 | HHsearch (proba=100.00%) |
| Portal | 4 | lcl|NC_016765.1_cdsid_YP_005098207.1 | HHsearch (proba=100.00%) |
| TermL | 2 | lcl|NC_016765.1_cdsid_YP_005098205.1 | Blast (e-value=0) |
| MTP | 16 | lcl|NC_016765.1_cdsid_YP_005098219.1 | Blast (e-value=2e-99) |
| Sheath | NOT FOUND | ||
| Ad1 | 9 | lcl|NC_016765.1_cdsid_YP_005098212.1 | HHsearch (proba=100.00%) |
| Hc1 | 10 | lcl|NC_016765.1_cdsid_YP_005098213.1 | HHsearch (proba=100.00%) |
| Ne1 | 14 | lcl|NC_016765.1_cdsid_YP_005098217.1 | HHsearch (proba=100.00%) |
| Tc1 | 15 | lcl|NC_016765.1_cdsid_YP_005098218.1 | HHsearch (proba=100.00%) |
| Ad2 | NOT FOUND | ||
| Hc2 | NOT FOUND | ||
| Tc2 | NOT FOUND | ||
| Ad3 | NOT FOUND | ||
| Hc3 | NOT FOUND | ||
| Ad4 | NOT FOUND |
| Proteins Superfamilies | Observed Intergene Distance | Expected Intergene Distance |
|---|---|---|
| Tc1<->Hc1 | 5 | 3 |
| Ne1<->Hc1 | 4 | 2 |
| Tc1<->Ad1 | 6 | 4 |
| Ne1<->Ad1 | 5 | 3 |
| Protein Superfamily | Corresponding Protein in Aclame | Aclame Phage | Seq. Identity (%) |
|---|---|---|---|
| MCP | protein:vir:4339 protein:vir:100135 protein:vir:97053 | D3 phi1026b OP1 | 99 % 60 % 47 % |
| Portal | protein:vir:4337 protein:vir:100150 protein:vir:81072 | D3 phi1026b Xop411 | 100 % 47 % 39 % |
| TermL | protein:vir:4335 protein:vir:481 protein:vir:4508 | D3 P27 V | 98 % 48 % 47 % |
| MTP | protein:vir:4349 protein:vir:1893 protein:vir:100241 | D3 HK022 Bcep176 | 99 % 62 % 23 % |
| Ad1 | protein:vir:4342 protein:vir:100245 protein:vir:81069 | D3 Bcep176 Xop411 | 97 % 25 % 23 % |
| Hc1 | protein:vir:4343 protein:vir:1890 protein:vir:81177 | D3 HK022 Geobacillus virus E2 | 97 % 41 % 28 % |
| Ne1 | protein:vir:4347 protein:vir:1891 protein:vir:105089 | D3 HK022 phiKO2 | 100 % 44 % 29 % |
| Tc1 | protein:vir:4348 protein:vir:1892 protein:vir:100242 | D3 HK022 Bcep176 | 100 % 58 % 25 % |